Paper: Lee, W.P., Hsiao, Y.T. and Hwang, W.C. (2014). Designing a parallel evolutionary algorithm for inferring gene networks on the cloud computing environment. BMC Systems Biology, 8:5.
Gene regulatory networks (GRNs) are like the instruction manuals for living cells. They show exactly how genes talk to each other, acting like switches to turn other genes on or off, which ultimately changes how a cell behaves. Figuring out the layout of these networks from biological data is a massive puzzle that is incredibly slow for computers to solve. This paper shows how we can use cloud computing to make this complex scientific problem much more manageable.
The problem with large gene networks
To understand these networks, the researchers use complex mathematical formulas (S-system model) to track how gene activity changes over time. The main challenge is finding the exact numbers (parameters) that make the math match the real-life data. As the number of genes grows, this becomes a giant guessing game.
Evolutionary algorithms are a natural fit for this kind of search: they explore the solution space in a somewhat random way and can avoid getting stuck in poor solutions. But they hit two walls when networks get large. First, the population tends to lose diversity too early and converges before finding a good answer. Second, computation time scales very badly.
The hybrid GA-PSO approach
To fix the variety problem, the authors combined two different computer methods into a hybrid called iGA-PSO, the idea is to get the best of both: PSO is good at local fine-tuning, GA at maintaining diversity through crossover and mutation. On top of that, they use a parallel island model: instead of keeping all the guesses in one big pool, they split them up into smaller, independent groups (islands). These groups work on the problem separately but occasionally share their best answers with neighboring islands. This keeps the ideas fresh and stops the program from getting stuck on a bad answer. Because each gene’s problem is distinct, it maps perfectly to its own island and its own computer.
Where cloud computing actually earns its place
The “island model” is a great idea, but running it on massive, dedicated supercomputers is too expensive for most scientists. The major breakthrough in this paper is making this process run on Hadoop MapReduce. MapReduce is a cloud computing setup that works on regular, affordable computer hardware.
Cloud computing is not just about raw speed here. It is about making certain classes of problems accessible to groups that cannot afford dedicated clusters.
MapReduce splits the work into map tasks and reduce tasks, with a master node coordinating everything and slave nodes doing the actual computation in parallel.
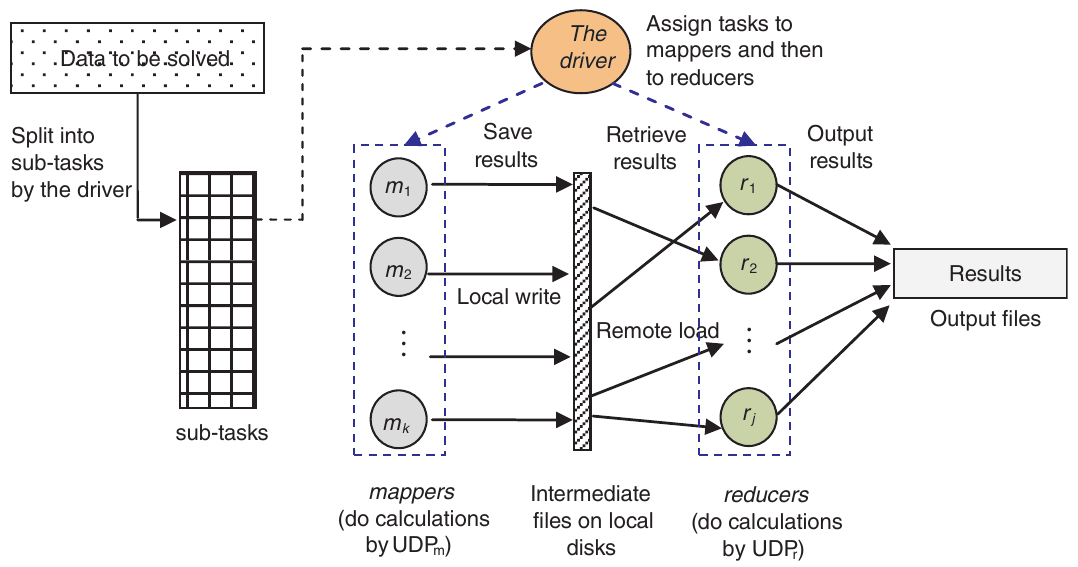
Figure 1: General workflow of a MapReduce process, with mappers handling independent subtasks and reducers combining the results.
The islands are arranged in a hypercube topology, so migration only happens between neighbouring nodes. Communication overhead stays low while good solutions still spread across the population over time.
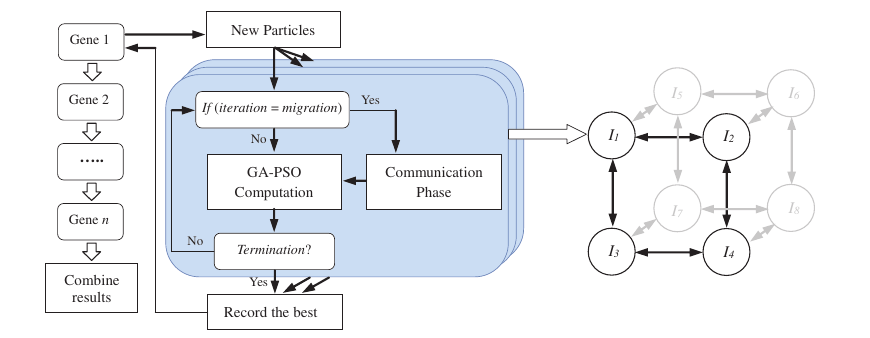
Figure 3: Parallel island model of the iGA-PSO approach, with the hypercube communication topology on the right.
Hadoop handles all the background work, like distributing the data and preventing crashes. Because dozens of cloud computers are sharing the workload, tracking down a massive 125-gene network becomes solvable in a reasonable amount of time.
Does it work?
Yes! The researchers tested their system using real data from yeast cells, specifically looking at networks of 25, 50, 100, and 125 genes. The cloud-based hybrid method (iGA-PSO) consistently beat the older, simpler methods. It found more accurate networks with fewer errors, and the cloud computers made the whole process significantly faster as the networks grew larger.
The takeaway: The individual pieces of this puzzle (Genetic Algorithms, Particle Swarms, and the island model) already existed. The authors’ real achievement was engineering a way to wire them all together on standard cloud software like MapReduce.
Real-life biology experiments in a lab are slow and costly. Having a fast, affordable, cloud-based computer tool to predict how genes interact is a massive benefit to the field.